About
 I am a Research Fellow at Museums Victoria, working on Jane Melville’s lab on how Quaternary changes affected past and recent reptile communities, using rich fossil data. I am also currently associated with the Australian National University (ANU), collaborating with the Moritz and Cardillo labs.
I am a Research Fellow at Museums Victoria, working on Jane Melville’s lab on how Quaternary changes affected past and recent reptile communities, using rich fossil data. I am also currently associated with the Australian National University (ANU), collaborating with the Moritz and Cardillo labs.
Previously, I worked as a data analyst, part of the Science & Decision Support team at the Atlas of Living Australia (CSIRO), a collaborative, digital, open infrastructure that pulls together Australian biodiversity data from multiple sources, making it accessible and reusable. I work in partnership with EcoCommons, a platform for digital modelling and analysis needs on ecological and environmental research. I also collaborated with the Australian National Wildlife Collection (ANWC – CSIRO), as a Relocation Technician.
I completed a postdoctoral position at the University of Michigan, at Lacey Knowles’ lab, interested in the species delimitation process, linking micro and macroevolution. I did my Ph.D. at ANU, at Craig Moritz’s lab. My dissertation concerned macroecological questions and landscape genomics related to lizards in savannas environments, mostly in Australia and Brazil. In my Master’s (University of Brasilia, Brazil), I worked with phylogenetic diversity and habitat loss in Neotropical pitvipers. I also assisted in the preparation of the redlist of lizards and amphisbaenians of Brazil.
I am a proud member of two Brazilian women in STEM initiatives, H2H (Herpetologia Segundo herpetólogas) and Rede Kunhã Asé, aiming to promote equity and value the role of Women in Science.
Any questions? Do you want to chat? Feel free to contact me by email
(jfenker@museum.vic.gov.au) and follow my updates on Twitter (@jehzim).
Current Research
Diversification of Lizards in savanna environments
Ecomorphological diversification

Tropical savannas are intermediate systems between dry xerophytic areas and moist deciduous forests. Their exhibit a wide variety of local environments, which allows a high number of species to coexist, with high levels of endemism. Both South America and Australia tropical savannas have highly diverse herpetofauna. Similar biogeographic events profoundly marked both continents-from a common Gondwanan origin, followed by a long period of isolation and recent colonization and diversity of lineages from the north. In face of the analogous histories of environments, what can we infer from the comparative biogeography, morphology, and diversity of these two squamates faunas? How does the morphospace volume differ between Cerrado and AMT squamate communities? What is the relationship between morphospace volume and evolutionary history and how does it differ between those regions? How similar these communities are compared to null expectations? And finally, can we observe convergence on squamate assemblages on Cerrado and AMT?
_______________________________________________________________________________________________________________
Evolutionary Conservation of the Cerrado

The Cerrado is a biodiversity hotspot and the most biodiverse tropical savanna in the world. Brazilian conservation policies often neglect non-forest ecosystems, and determining where the diversity of Cerrado taxa is concentrated is important for prioritizing conservation efforts. Combining diversity and endemism measures with an integrative approach using taxonomic, phylogenetic, and functional data we aimed to highlight key areas for the past and future maintenance of biodiversity in the Cerrado. We use refined phylogeographic and population-level lizards datasets to improve phylogenetic diversity estimates and an independently named taxa approach to support conservation planning for biota in areas where the taxonomy is incomplete. This work is published in Journal of Biogeography, presented at the Journal of Biogeography ECR feature blog, and won an award as “Top Cited article 2020-2021” by the journal!
_______________________________________________________________________________________________________________
Phylogeography and incipient speciation in Diporiphora agamids
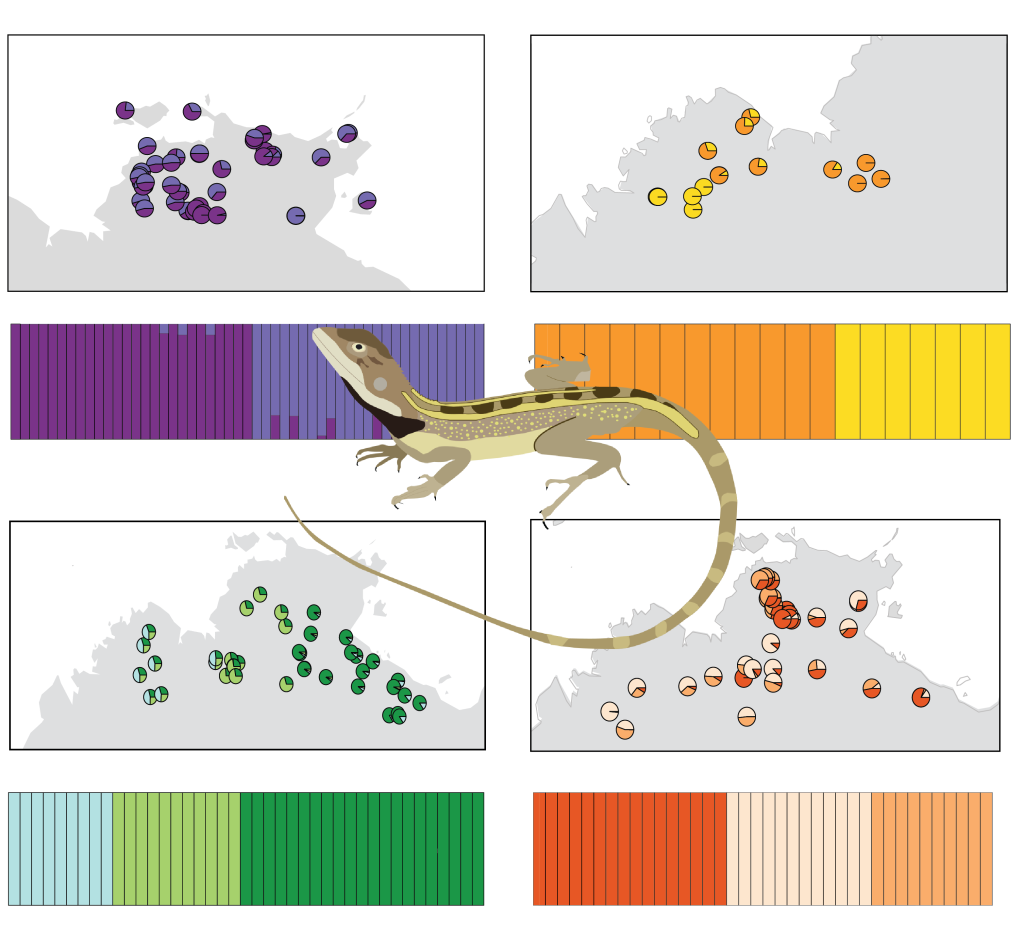
The Australian Monsoonal Tropics (AMT) is a vast tropical savanna with high levels of richness and endemism, associated with its environmental heterogeneity. Despite our growing knowledge surrounding species diversity, species have different ecological requirements and can respond differently to the same geographic events. The two-lined dragons (genus Diporiphora) are a diverse group of small diurnal lizards, relatively common in Australia. Taxonomic problems related to conservative morphological characteristics have long been recognized for the genus. Using a broad geographic and genomic sampling of mtDNA and SNP variation in Diporiphora dragons from across the AMT, we aimed to evaluate the impact of biogeographic breaks and climatic fluctuations on phylogeographic structuring and demographic history of species with diverse ecological requirements.
_______________________________________________________________________________________________________________
Predictors of Phylogeographic Structure and Landscape Genetics

Species dispersal rates vary depending on intrinsic movement rates and species habitat preference and are predicted to influence rates of speciation and extinction and hence spatial patterns of diversity. With a combination of population genetic summary statistics and landscape genomic analyses, based on mtDNA and extensive SNP datasets from geckos, dragons, and scincid lizards across the diverse and heterogeneous savanna-like Australian Monsoonal Tropics, we aim to explore how the environment and landscape features affect the gene flow of species across genera and with different habitat specialization across the same geographical region. We asked 1) does habitat use predict how landscape features influence the degree of gene flow across a common environment? and 2) how migration and population expansion affect species with different habitat use? Then, we explored predictors of the scale and depth of the phylogeographic structure, testing whether microevolutionary processes explain differences in the geographic scale and depth of phylogeographic lineages. This work is published in Molecular Ecology.
_______________________________________________________________________________________________________________
